This is a collection of frequently asked questions about the Segmented Inner Organs of the Visible Human (SIO).
1 Is this the same as the Visible Human Project?
No, but it’s based on the Visible Human Project. The original color images of the Visible Human Male were reduced in size and geometrically aligned with the CT images by 3D registration, resulting in an isotropic, multi-spectral image volume. In a subsequent 3D segmentation, the anatomical components were identified and marked with object labels.
2 What is the resolution of the model?
The limiting factor is the voxel size of 1 mm x 1 mm x 1 mm. Structures smaller than that may only be represented as slight changes in color or CT intensity.
3 How accurate is the segmentation?
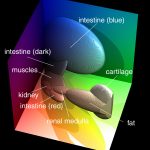 The segmentation was done with an advanced classifier in RGB color space, which can locate the object borders very precisely (see references).
The segmentation was done with an advanced classifier in RGB color space, which can locate the object borders very precisely (see references).
The VOXEL-MAN 3D-Navigator: Inner Organs atlas that is based on this very model was positively reviewed by several independent expert anatomists and radiologists.
4 Does the model include the muscles?
Yes, the model includes almost all of the muscle tissue of the torso. However, it is mostly labeled as “unclassified muscles”, only a few muscles of the abdominal region are explicitly labeled. Please have a look at the object list.
5 Does this model include all parts of the Inner Organs anatomical atlas?
Almost, but some thin arteries, nerves and lymph nodes that are not visible on the cross-sectional images were modeled with a tube editor and represented as polygonal surface meshes. They are not part of this model.
6 How can I get the model?
The model is available for research purposes and requires a signed license agreement. Please follow the instructions given on the Segmented Inner Organs page.
7 Can I have a look at the data first?
Sure, please download the sample file. This ZIP archive contains a set of 5 consecutive slices, each with color, CT, and label images. An object list is also included, which links the numbers in the label images to their anatomical meanings.
8 How is the label volume organized?
Just like color and CT data, the label volume comes as a sequence of numbered cross-sectional images. Color, CT and label images with the same image number represent different properties of the same voxels.
Instead of colors or intensity values, a label image contains numerical object labels. Each object is represented by at least one object number (e.g. ascending aorta 511, arch of aorta 512, descending aorta 513, abdominal aorta 514). Wherever you find an object number, the corresponding object is present. A complete list of object numbers can be found in the object list.
9 Why isn’t there anything to see on the label images?
These are 16 bit TIFF images, which some programs cannot display properly. Programs that can handle these data include ImageJ (free) or Adobe Photoshop. If the image still appears all black, adjust the brightness/contrast or level/window settings, respectively.
10 How can I create a volume dataset from these images?
The images need to be stacked on top of each other. For example, in ImageJ, use File > Import > Image Sequence … to create three image volumes from the color, CT, and label images, respectively.
11 Is this a polygonal surface model?
No, the data comes as a three-dimensional voxel model. If you need a polygon mesh, you can create it for yourself, using methods such as Marching Cubes.
12 Can I get the model as a 3DS, FBX, STL, OBJ or 3MF file?
All of these are file formats for computer-aided design, 3D graphics or 3D printing that contain surface representations, see previous question.
13 Can I use the SIO for FEM simulation?
Sure. However, as with the polygon models, you need to create your own finite element model from the voxel model.
14 Can I get the segmentation software?
The interactive segmentation that we use for all our virtual 3D models is implemented in the VOXEL-MAN My Cases simulator application. Note however that this version is optimized for radiological images and does not include the RGB classifier.
15 This sounds way too technical for me. How can I use this model to study human anatomy?
The VOXEL-MAN 3D-Navigator: Inner Organs interactive atlas of anatomy and radiology is available for free download under a Creative Commons license. It is based on this 3D model.
Back to the Segmented Inner Organs of the Visible Human